3.18 Pipes and Plots
REVIEW A common scenario for pipeline processing is to prepare data for plotting. Indeed, plotting itself has a pipeline type concept where we build a plot by adding layers to it.
Below the rattle::weatherAUS dataset is
dplyr::filter()ed for observations from four Australian
cities. We dplyr::filter() observations that have missing
values for the variable Temp3pm using an embedded
pipeline. The embedded pipeline pipes the Temp3pm data
through the base::is.na() function which tests if the value
is missing. These results are then piped to magrittr::not()
which inverts the true/false values so that we include those that are
not missing.
A plot is generated using ggplot2::ggplot() into which we pipe the processed dataset. We add a geometric layer using ggplot2::geom_density() which consists of a density plot with transparency specified through the argument. We also add a title and label the axes using ggplot2::labs().
cities <- c("Canberra", "Darwin", "Melbourne", "Sydney")
ds %>%
filter(location %in% cities) %>%
filter(temp_3pm %>% is.na() %>% not()) %>%
ggplot(aes(x=temp_3pm, colour=location, fill=location)) +
geom_density(alpha=0.55) +
labs(title = "Density Distributions of the 3pm Temperature",
x = "Temperature Recorded at 3pm",
y = "Density")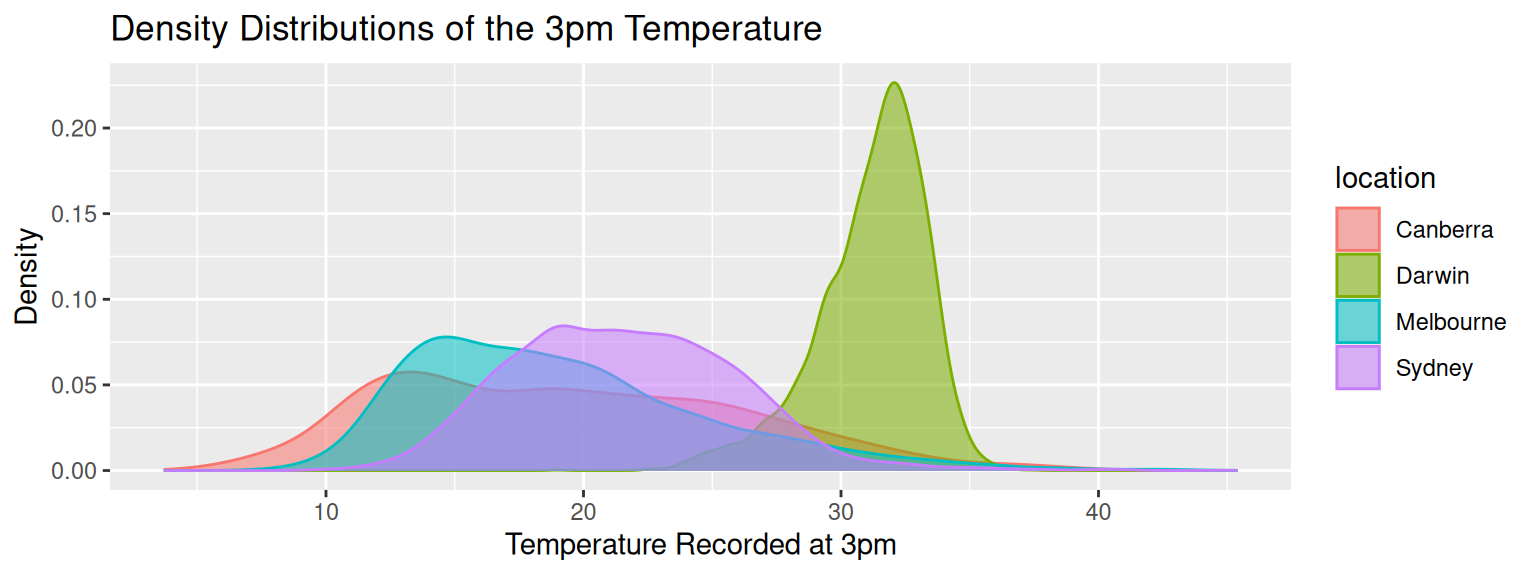
We now observe and tell a story from the plot. Our narrative will begin with the observation that Darwin has quite a different and warmer pattern of temperatures at 3pm than Canberra, Melbourne and Sydney. Canberra is on the colder side with Sydney generally warmer than Melbourne!
Your donation will support ongoing availability and give you access to the PDF version of this book. Desktop Survival Guides include Data Science, GNU/Linux, and MLHub. Books available on Amazon include Data Mining with Rattle and Essentials of Data Science. Popular open source software includes rattle, wajig, and mlhub. Hosted by Togaware, a pioneer of free and open source software since 1984. Copyright © 1995-2022 Graham.Williams@togaware.com Creative Commons Attribution-ShareAlike 4.0
